- Documentation
- Community
- News
-
API
- pyhrf.boldsynth package
- pyhrf.jde package
- Subpackages
- Submodules
- pyhrf.jde.asl module
- pyhrf.jde.asl_2steps module
- pyhrf.jde.asl_physio module
- pyhrf.jde.asl_physio_1step module
- pyhrf.jde.asl_physio_1step_params module
- pyhrf.jde.asl_physio_det_fwdm module
- pyhrf.jde.asl_physio_hierarchical module
- pyhrf.jde.asl_physio_joint module
- pyhrf.jde.beta module
- pyhrf.jde.drift module
- pyhrf.jde.hrf module
- pyhrf.jde.jde_multi_sess module
- pyhrf.jde.jde_multi_sujets module
- pyhrf.jde.jde_multi_sujets_alpha module
- pyhrf.jde.models module
- pyhrf.jde.noise module
- pyhrf.jde.samplerbase module
- pyhrf.jde.wsampler module
- pyhrf.sandbox package
- pyhrf.stats package
- pyhrf.test package
- Submodules
- pyhrf.test.analysertest module
- pyhrf.test.boldsynthTest module
- pyhrf.test.commandTest module
- pyhrf.test.core_test module
- pyhrf.test.graphtest module
- pyhrf.test.iotest module
- pyhrf.test.jdetest module
- pyhrf.test.rfir_test module
- pyhrf.test.seppotest module
- pyhrf.test.statsTest module
- pyhrf.test.test module
- pyhrf.test.test_glm module
- pyhrf.test.test_jde_multi_subj module
- pyhrf.test.test_jde_vem_asl module
- pyhrf.test.test_jde_vem_bold module
- pyhrf.test.test_jde_vem_tools module
- pyhrf.test.test_jde_vem_tools_UtilsC module
- pyhrf.test.test_jde_vem_tools_asl module
- pyhrf.test.test_ndarray module
- pyhrf.test.test_paradigm module
- pyhrf.test.test_parallel module
- pyhrf.test.test_parcellation module
- pyhrf.test.test_plot module
- pyhrf.test.test_rfir module
- pyhrf.test.test_sampler module
- pyhrf.test.test_sandbox_physio module
- pyhrf.test.test_treatment module
- pyhrf.test.test_xml module
- pyhrf.test.toolsTest module
- Submodules
- pyhrf.tools package
- pyhrf.ui package
- pyhrf.validation package
- Submodules
- pyhrf.validation.config module
- pyhrf.validation.valid_beta_estim module
- pyhrf.validation.valid_jde_asl module
- pyhrf.validation.valid_jde_asl_physio module
- pyhrf.validation.valid_jde_asl_physio_alpha module
- pyhrf.validation.valid_jde_bold_mono_subj_multi_sess module
- pyhrf.validation.valid_jde_bold_mono_subj_sess module
- pyhrf.validation.valid_jde_vem_asl module
- pyhrf.validation.valid_rfir module
- pyhrf.validation.valid_rndm_field module
- pyhrf.validation.valid_sandbox_parcellation module
- Submodules
- pyhrf.vbjde package
- pyhrf.xmliobak package
- pyhrf.configuration module
- pyhrf.core module
- pyhrf.glm module
- pyhrf.graph module
- pyhrf.grid module
- pyhrf.ndarray module
- pyhrf.paradigm module
- pyhrf.parallel module
- pyhrf.parcellation module
- pyhrf.plot module
- pyhrf.rfir module
- pyhrf.surface module
- pyhrf.usemode module
- pyhrf.version module
- pyhrf.xmlio module


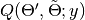 at a given iteration
at a given iteration